Code
ggplot(data = data_lionfish,
mapping = aes(x = total_length_mm,
y = total_weight_gr)) +
geom_point() +
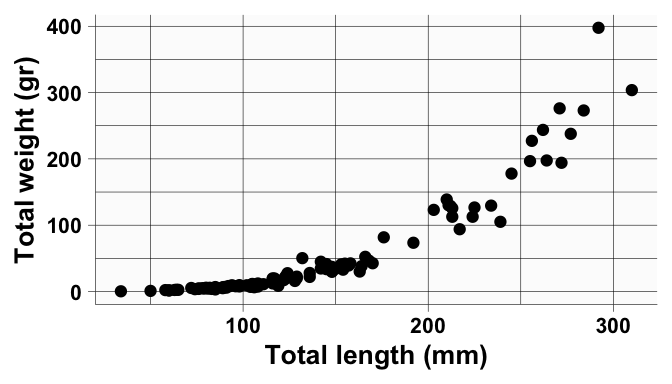
labs(x = "Total length (mm)",
y = "Total weight (gr)") +
theme(axis.line = element_line(color = "black",
linewidth = 0.5),
axis.ticks = element_line(color = "black",
linewidth = 0.5),
panel.grid.major = element_line(color = "black",
linewidth = 0.5),
panel.grid.minor = element_line(color = "black",
linewidth = 0.5),
panel.background = element_rect(fill = "lightblue"),
axis.text = element_text(color = "black"),
text = element_text(color = "black",
face = "bold",
size = 12))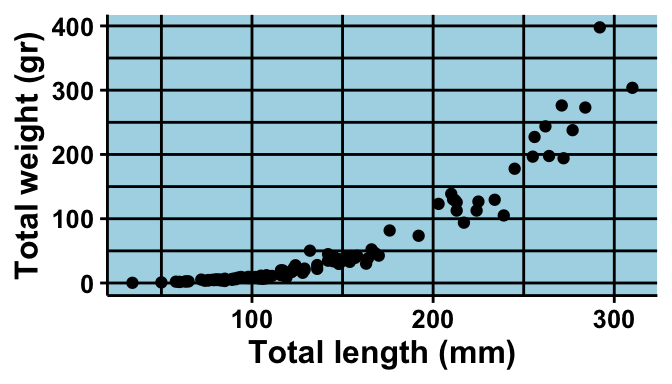
Code
ggplot(data = data_lionfish,
mapping = aes(x = total_length_mm)) +
geom_histogram() +
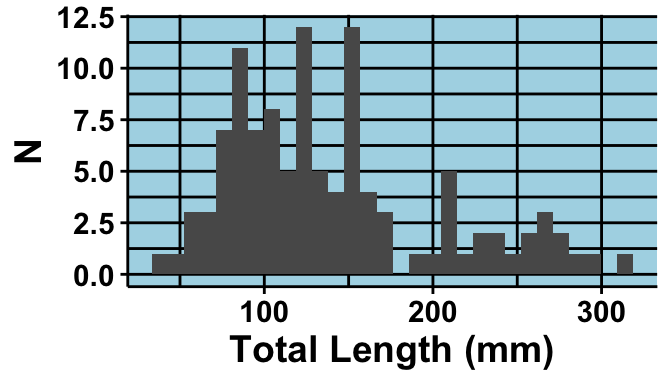
labs(x = "Total Length (mm)",
y = "N") +
theme(axis.line = element_line(color = "black",
linewidth = 0.5),
axis.ticks = element_line(color = "black",
linewidth = 0.5),
panel.grid.major = element_line(color = "black",
linewidth = 0.5),
panel.grid.minor = element_line(color = "black",
linewidth = 0.5),
panel.background = element_rect(fill = "lightblue"),
axis.text = element_text(color = "black"),
text = element_text(color = "black",
face = "bold",
size = 14))