Working with spatial data in R
EVR 628- Intro to Environmental Data Science
Rosenstiel School of Marine, Atmospheric, and Earth Science and Institute for Data Science and Computing
Announcements
Upcoming EVR Seminar

Last Week
- Spatial data can be represented in a
vectormodel, or arastermodel - We focused on
vectordata, which is represented by points, lines, polygons (and combination of these) vectordata are often saved as shapefiles, geoJSON files, or geopackage files- In
R, we use thesimple featuresstandard to work with vector data via{sf} - Regardless of the file extension (
.shp,.gpkg…), we read them withsf::read_sf() sfobjects have three main components:- CRS: Where is the origin
- geometry: The spatial portion of the data
- attributes: The characteristics associated to each point, line, or polygon
Learning Objectives
By the end of this week, you should be able to
- Identify whether a map was built with
rasterorvectordata (or both) - Load and work with
rasterobjects with{terra} - Visualize and manipulate
rasterdata with{tidyterra}
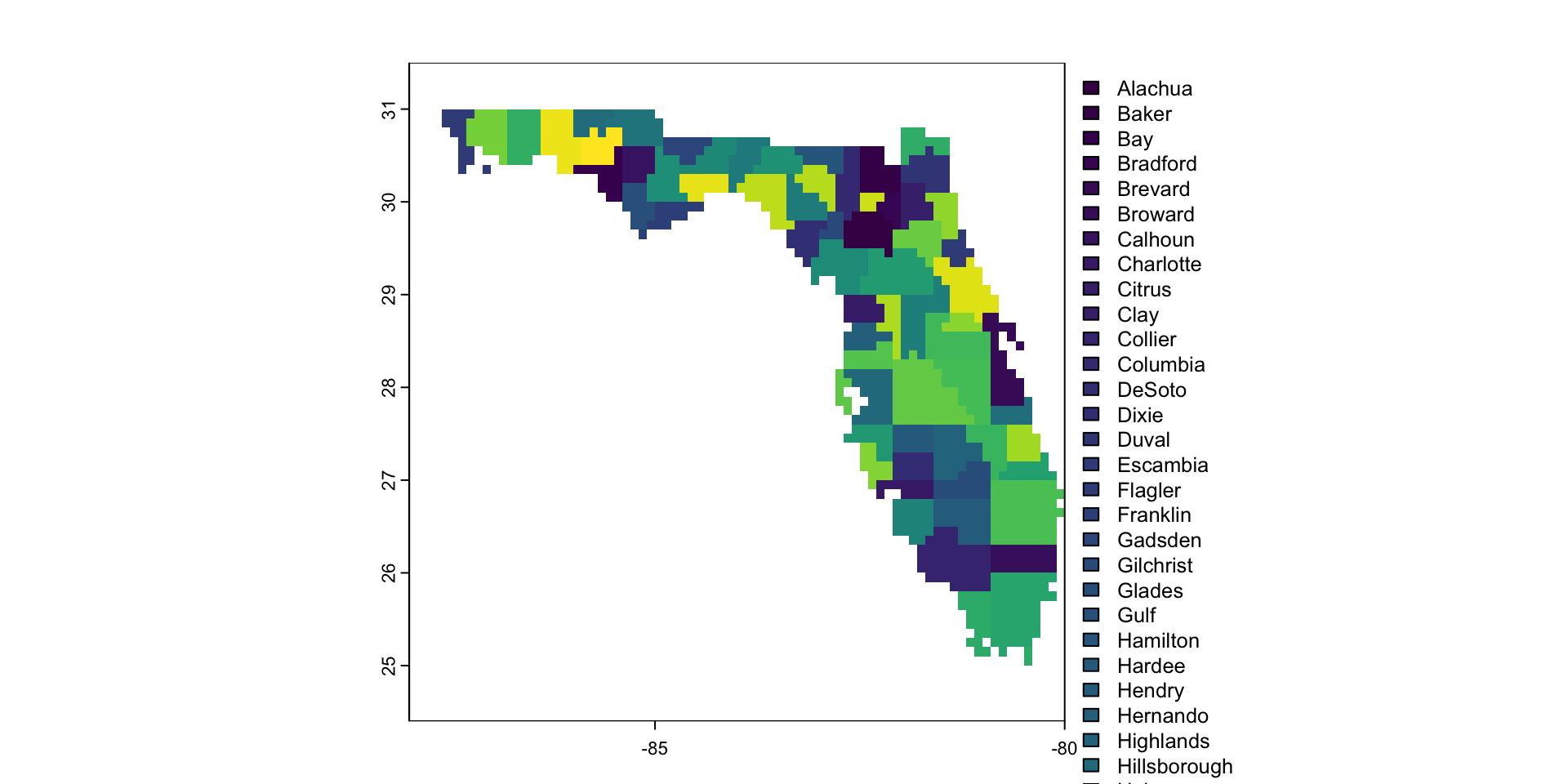
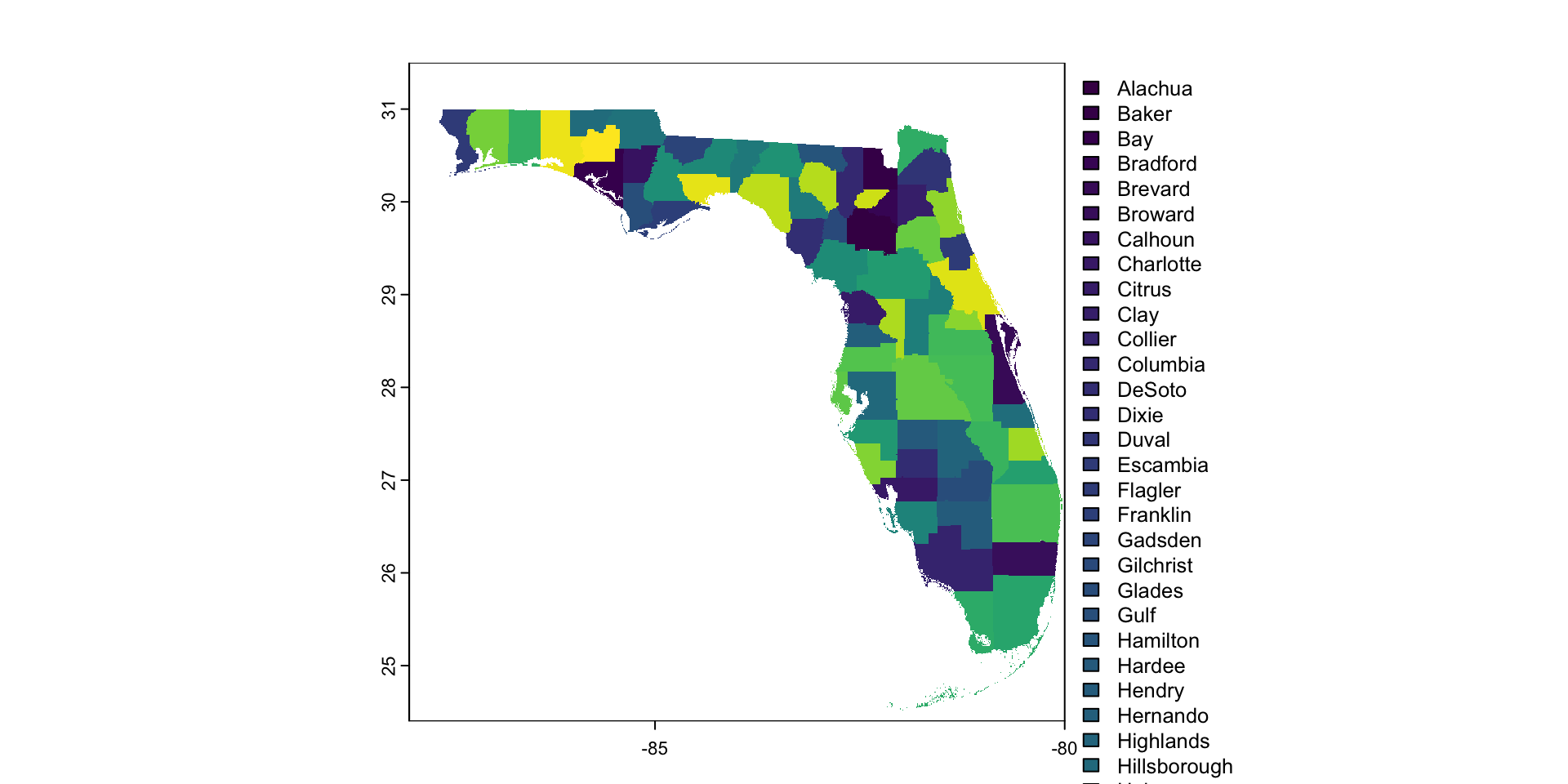
Raster Data Model
Raster Data Model
- The world is represented with a continuous grid
- Grid is made up of grid cells or pixels
- All pixels are the same size1
- For most applications, grid cells will be “square”
Raster Data Model
Raster Data Model
Common Applications:
- Usually used to represent continuous information:
- Elevation / Depth
- Temperature
- Population density
- Spectral data (in remote sensing applications)
- Discrete data can also be used
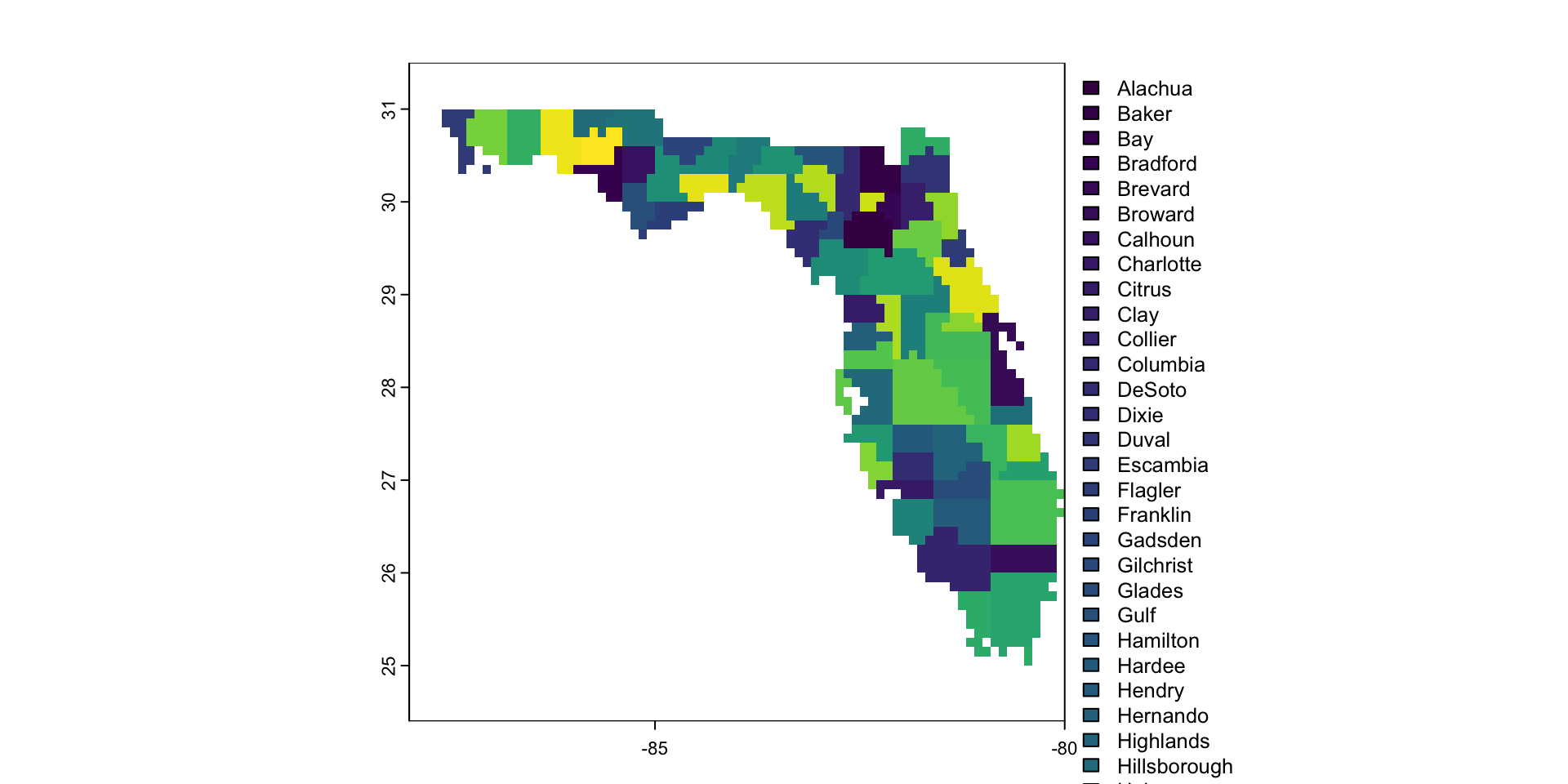
Raster Data Model
Common extensions include:
- .tiff / .tif:
- “Tagged Image File Format”
- .asc
- .nc: Common in remote sensing and climate model output
TIFF or “GeoTiff” is simply an image made of pixels
Components of a Raster Object
- Two main components:
- Header: metadata with CRS, extent, and origin
- Matrix: the data we want to represent
- x-coordinates = columns
- y-coordinates = rows
- In
terrathe “origin” of the matrix is the top-left corner
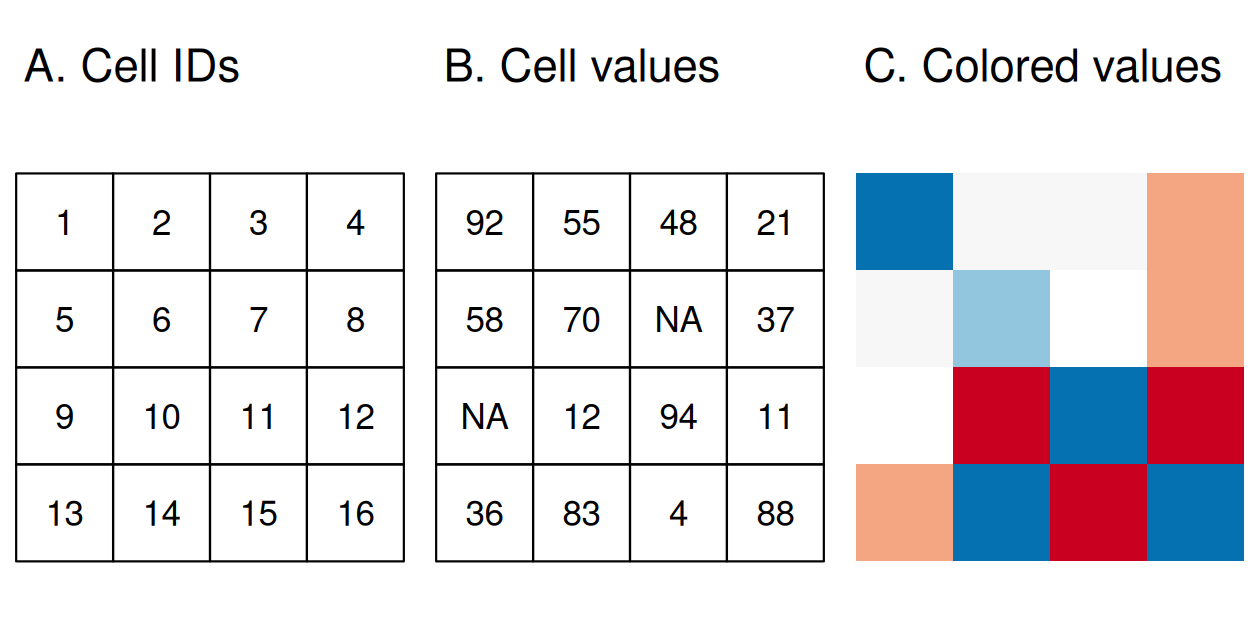
Raster Is Faster. . . But Why?
- There is a fundamental relationship between resolution, extent, and origin:
\[ \mathrm{resolution_x} = \frac{x_{max} - x_{min}}{n_{col}} \]
- Raster data are typically “lighter” (and faster to work with) because we don’t need to store the coordinates for all corners of each cell
Q: How many coordinates does R need to store if you specify the origin, extent, and resolution?
A: One coordinate pair, lattitude and longitude for the origin
- In contrast to vector data, the cell of one raster layer can only hold a single value
Raster Is Faster. . . But Why?

- I just need to store the coordinates for the center of cell #1
- Coordinates for cell number 2 are \((X_1 + resolution, Y_1)\)
Raster Data in R
The terra Package
The terra Package: Example
- I can read a raster file into using the function
rast()
class : SpatRaster
size : 3600, 7200, 1 (nrow, ncol, nlyr)
resolution : 0.05, 0.05 (x, y)
extent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)
coord. ref. : lon/lat WGS 84 (EPSG:4326)
source : depth_raster.tif
name : depth_m
min value : -10273.276367
max value : 0.985951 - Q: What is the resolution of this raster?
- A: 0.05°
- Q: What is the extent?
- A: The world
Find info contained in the header with specific functions:
SpatExtent : -180, 180, -90, 90 (xmin, xmax, ymin, ymax)[1] 0.05 0.05[1] 3600 7200 1[1] 25920000[1] "depth_m"Visualizing Rasters in R
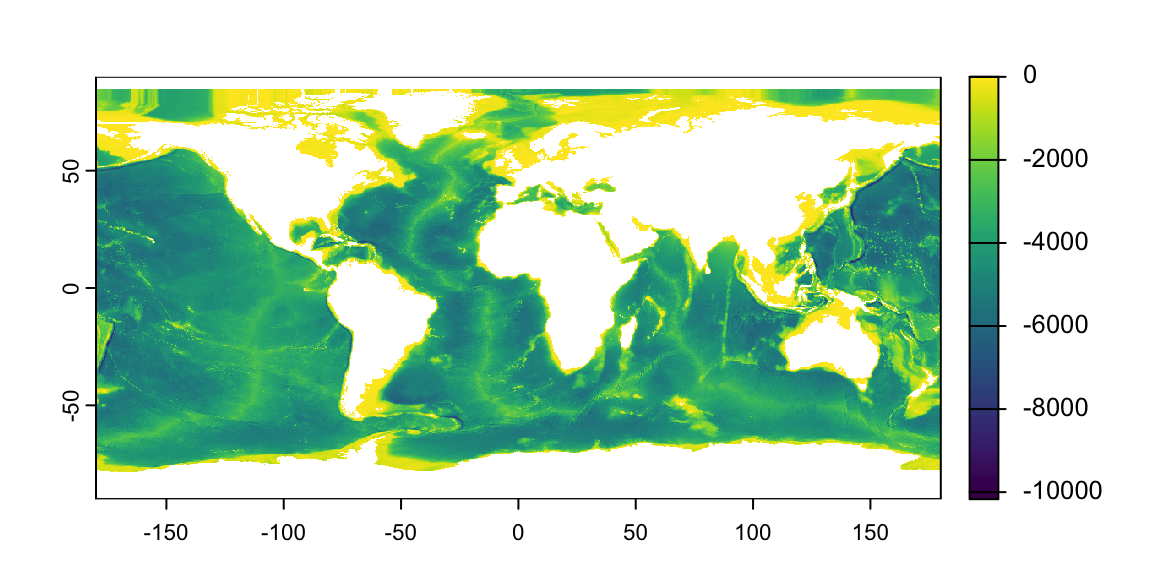
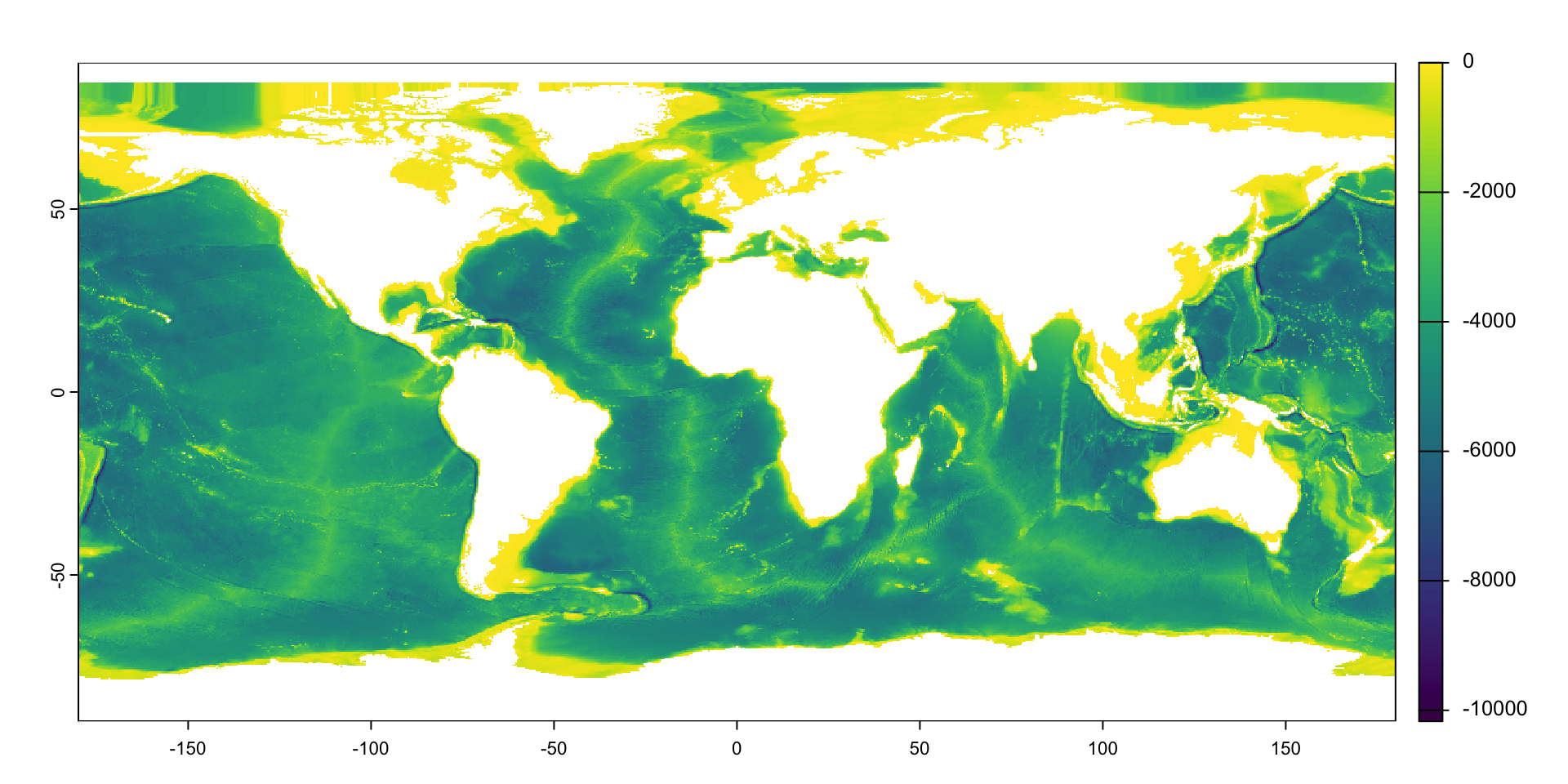
The fastest way is to use the base plot() function
Visualizing Rasters in R
As with vector data, I can layer different pieces of my plot
- Q: How are missing values represented?
- A: As transparent (white) pixels
Manipulating Raster Objects
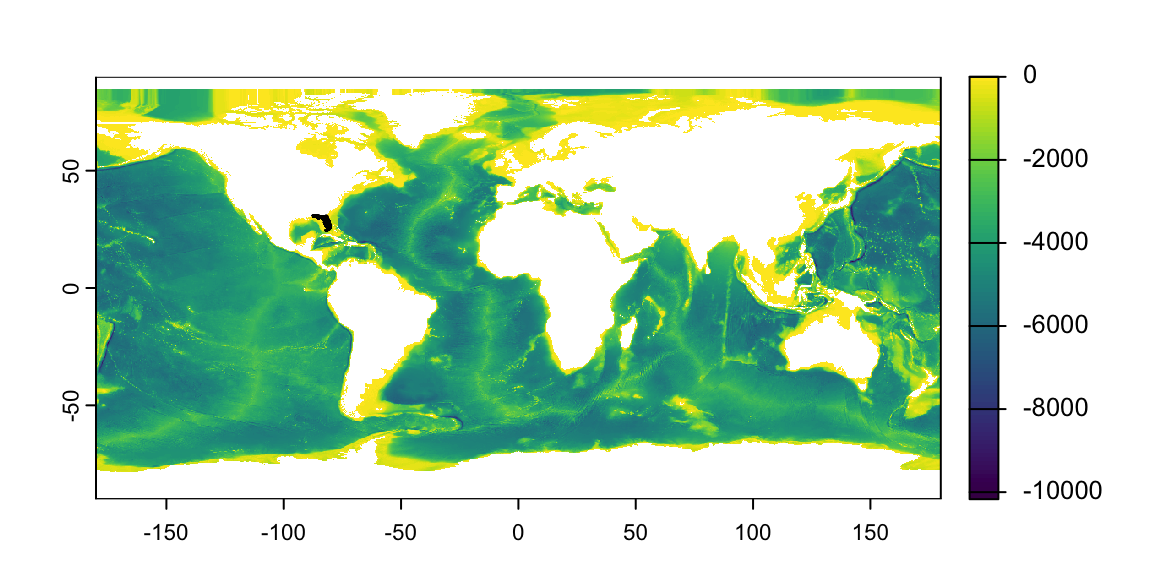
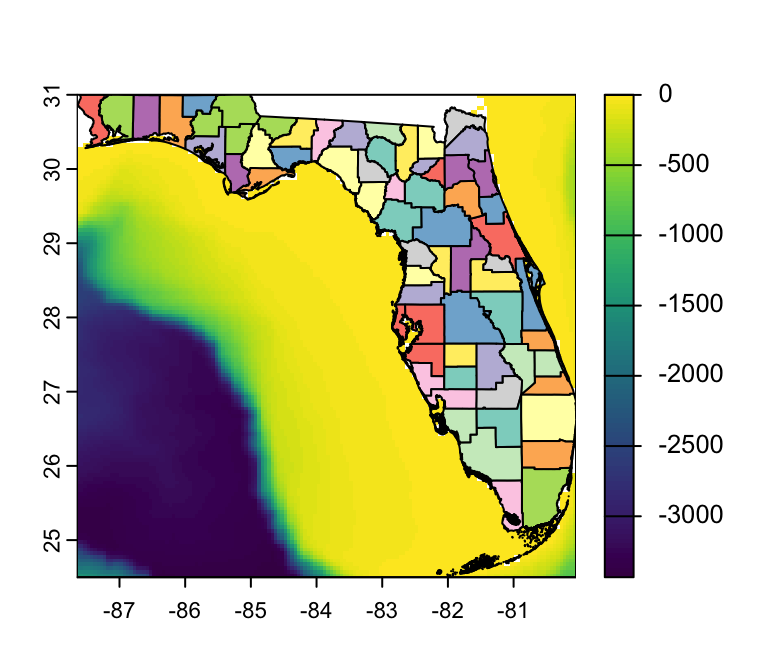
Task: Produce a map of water depth around FL
- Q: In human language, what do I have to do?
- A: “Zoom-in” on my global raster:
crop()1
Manipulating Raster Objects
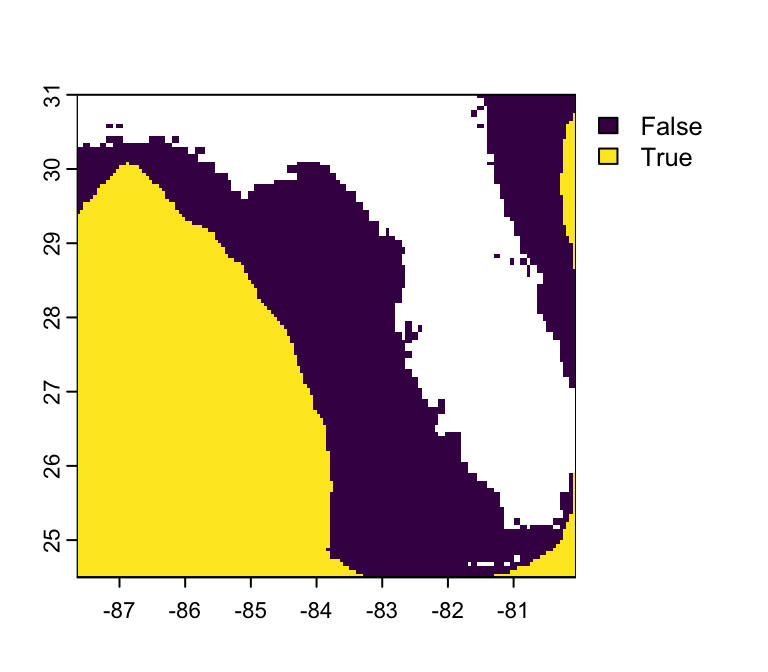
Task: Build the same map, but only show shallow (depth > -100 m) waters
- Rasters are just matrices, so we can use our trusty
binaryoperators
Manipulating Raster Objects
Task: Build the same map, but only show shallow (depth > -100 m) waters
- In a data.frame (or sf) object, you can remove rows (observations) based on values
- But a raster is not a tidy table. It’s a matrix, so removing rows wouldn’t be wise
- Instead of filtering data, we turn them into
NA
Tidy rasters in R: {tidyterra}
Tidy Rasters in R: {tidyterra}

{tidyterra}: Common methods of the tidyverse for objects created with {terra}
The {tidyterra} package:
- Developed by Diego Hernangómez (Hernangómez (2023))
- Extends, but doesn’t replace, the
terrapackage - Provides functions to manipulate and visualize the attribute portion of the raster
- Brings ggplot capabilities (
geom_spatraster())
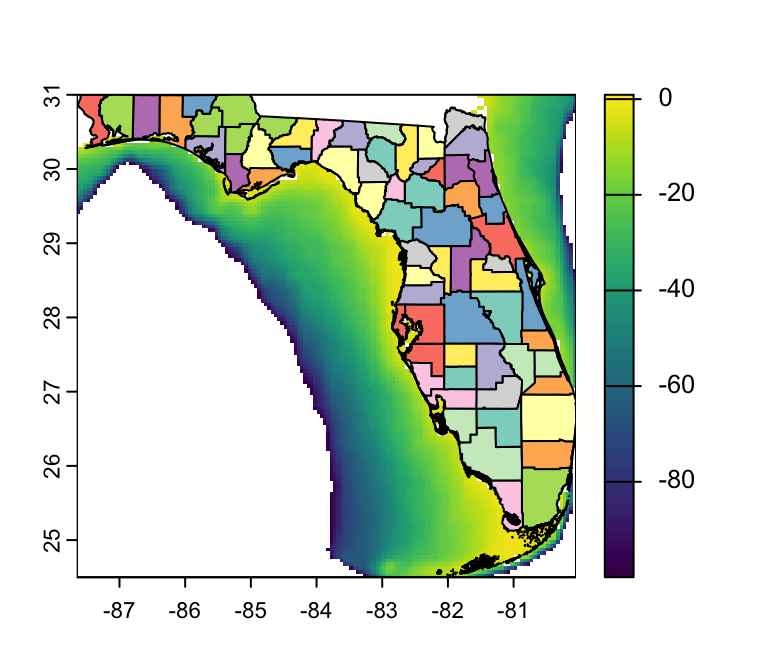
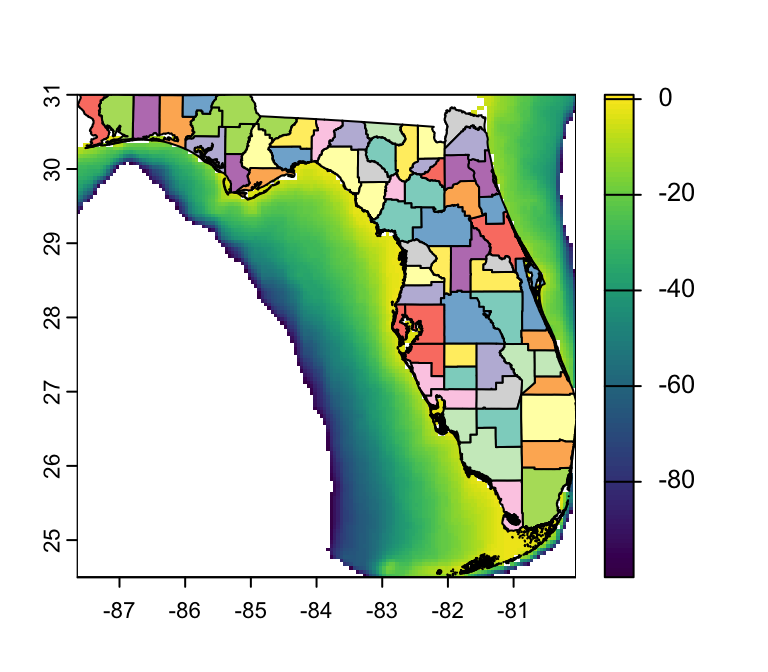
Visualizing Rasters in R: {tidyterra}
library(tidyterra) # Load terra
ggplot() + # Begin a ggplot
geom_spatraster(data = depth, # specify object to plot
aes(fill = depth_m)) + # specify layer to plot
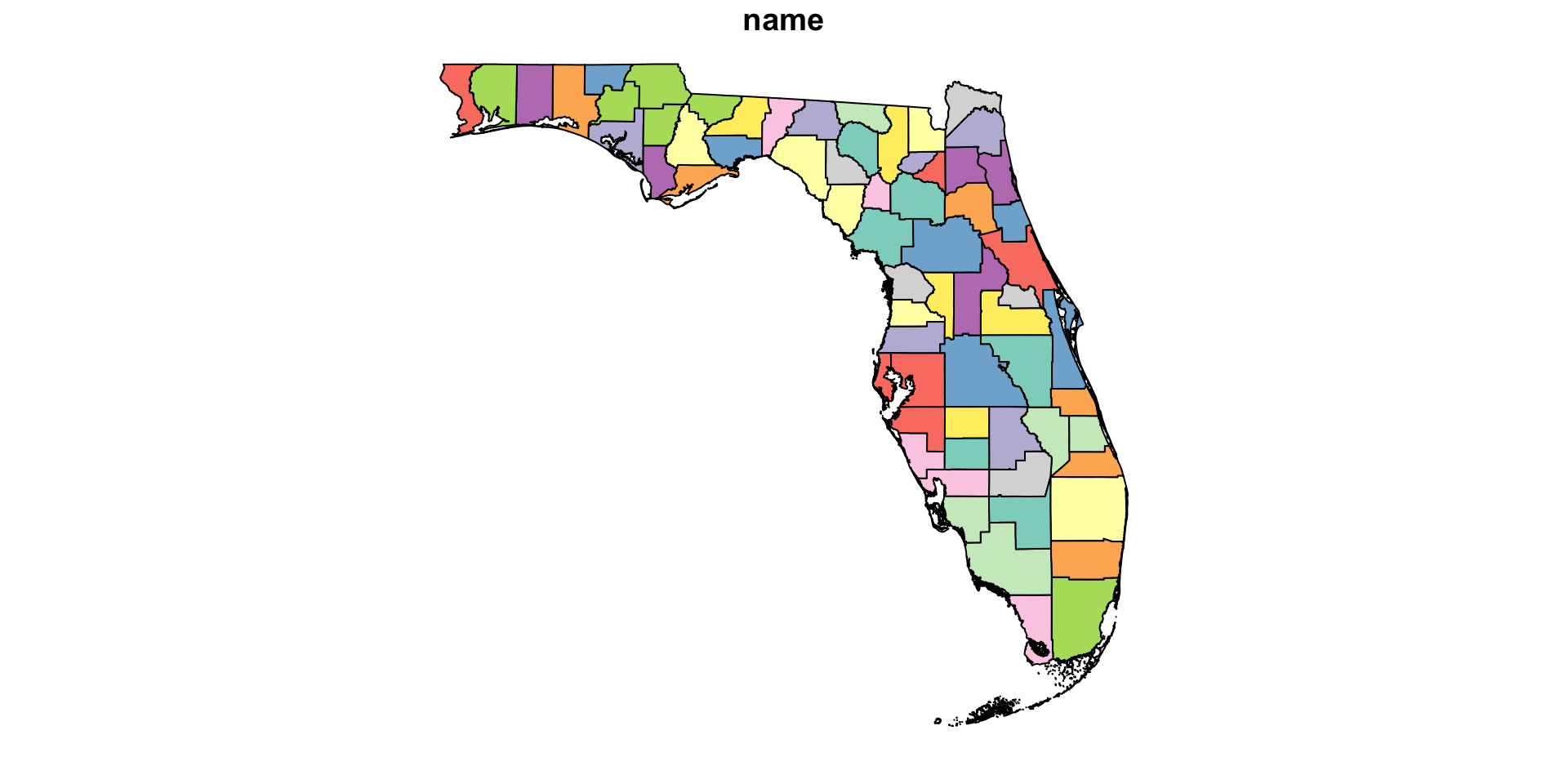
geom_sf(data = FL_counties) # Add FL counties on top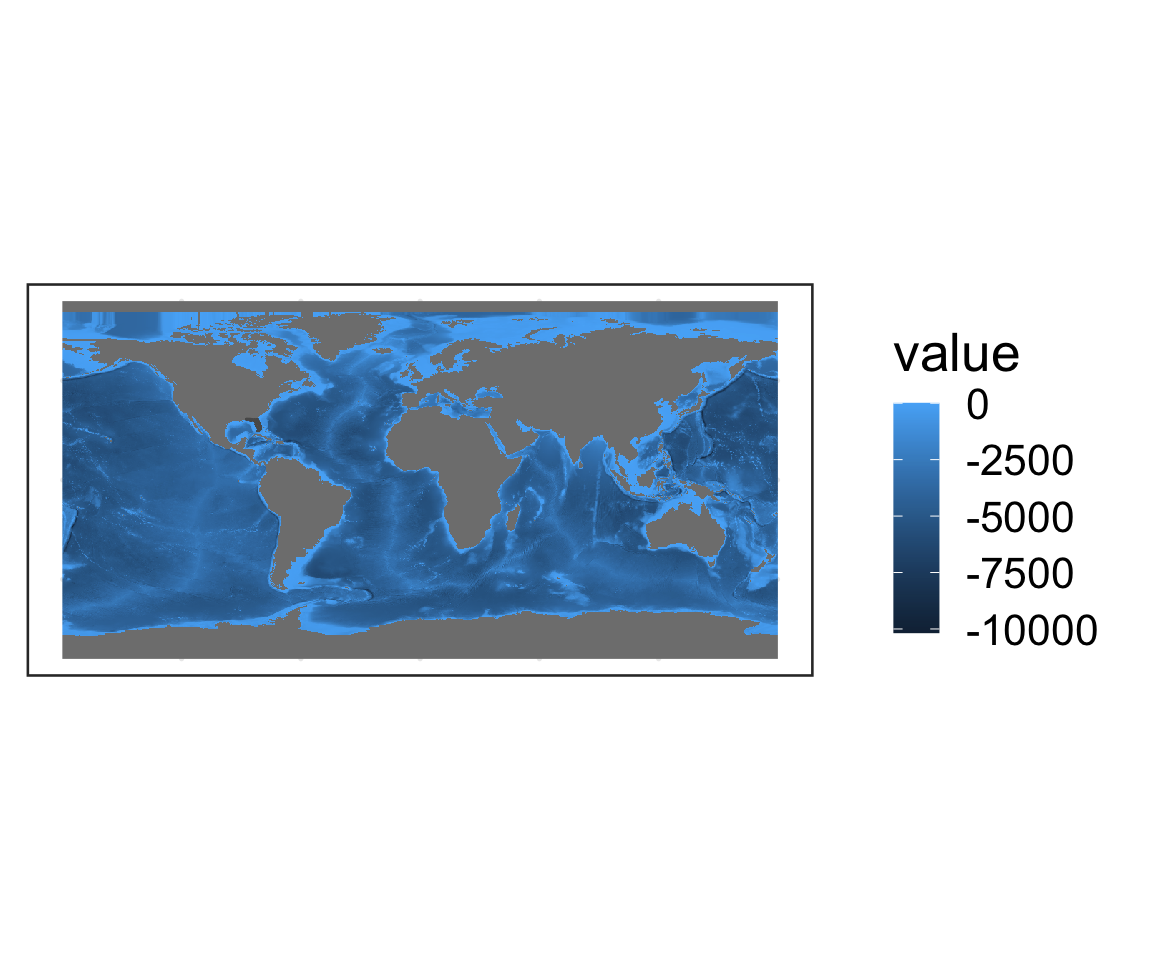
- Q: How are missing values represented?
- A: As gray pixels
Manipulating Rasters with tidyterra
Rasters with more than one attribute (layer)
Combining Rasters in terra
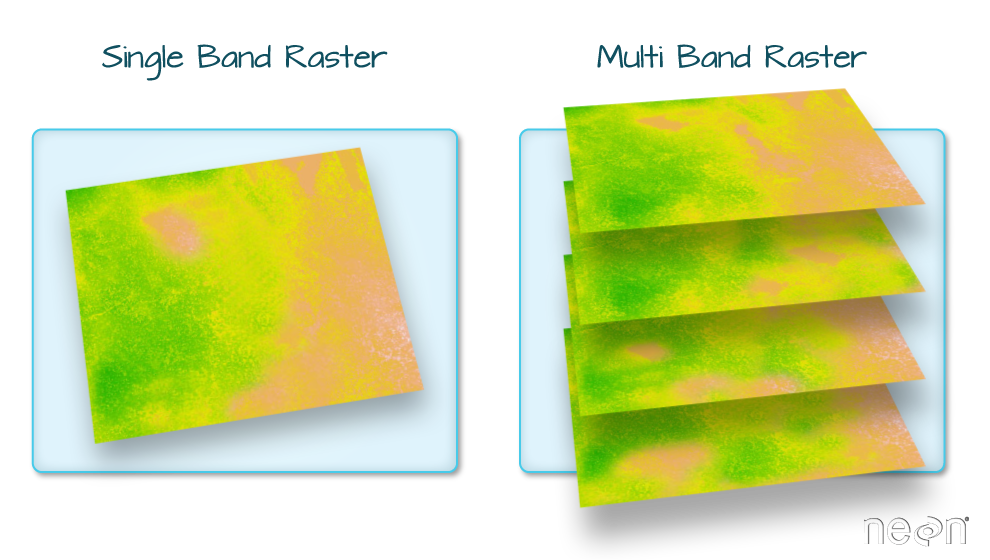

All layers must have the same extent and resolution
Combining Rasters in terra
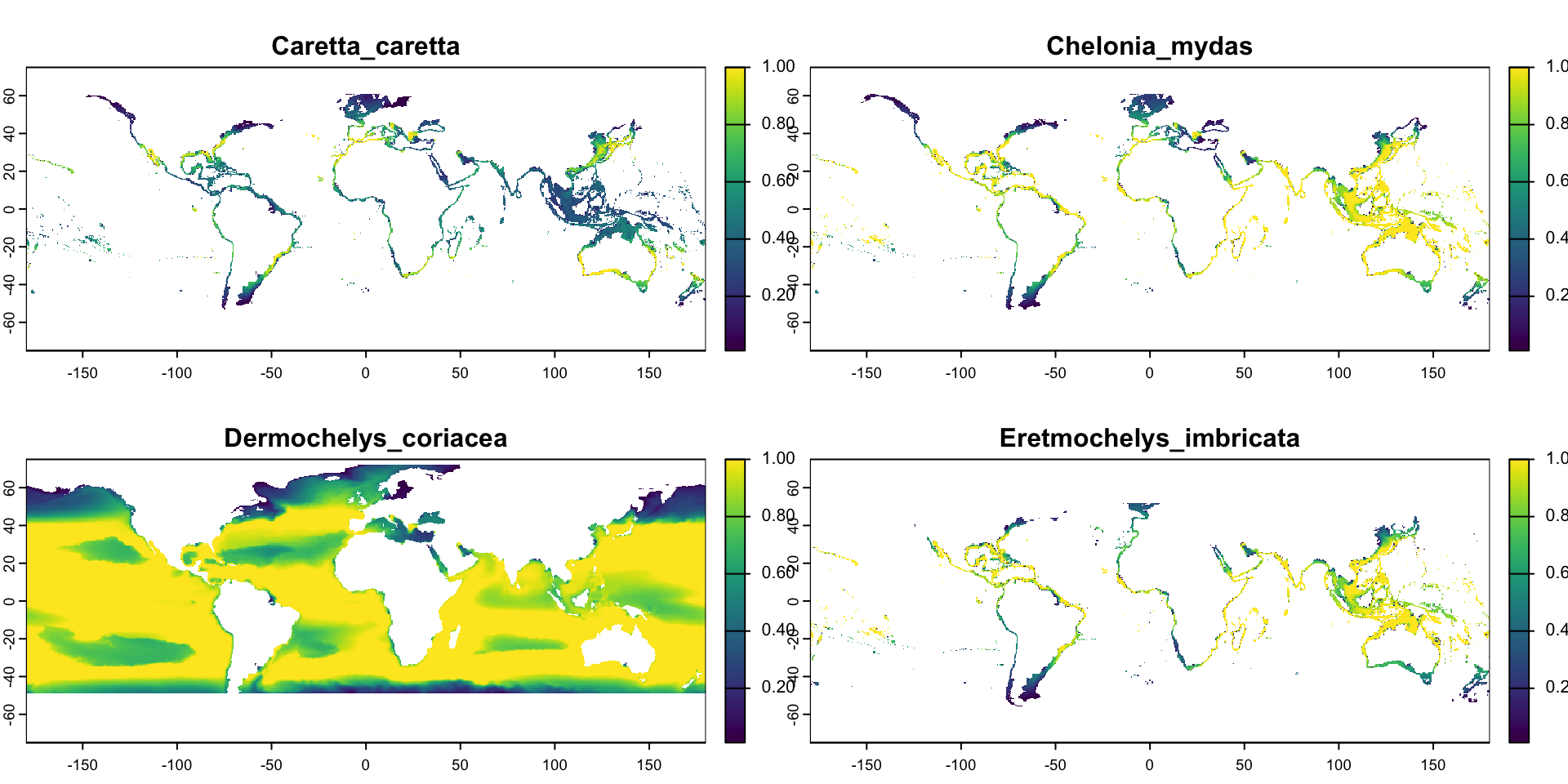
# Let's read habitat suitability models for four sea turtle species
log <- rast("data/raw/AquaMaps/Caretta_caretta.tiff")
green <- rast("data/raw/AquaMaps/Chelonia_mydas.tiff")
leather <- rast("data/raw/AquaMaps/Dermochelys_coriacea.tiff")
hawk <- rast("data/raw/AquaMaps/Eretmochelys_imbricata.tiff")
# We can combine them together using the `c()` function
turtles <- c(log, green, leather, hawk)
turtlesclass : SpatRaster
size : 300, 720, 4 (nrow, ncol, nlyr)
resolution : 0.5, 0.5 (x, y)
extent : -180, 180, -75, 75 (xmin, xmax, ymin, ymax)
coord. ref. : lon/lat WGS 84 (EPSG:4326)
sources : Caretta_caretta.tiff
Chelonia_mydas.tiff
Dermochelys_coriacea.tiff
Eretmochelys_imbricata.tiff
names : Caretta_caretta, Chelonia_mydas, Dermoch~oriacea, Eretmoc~bricata
min values : 0.01, 0.01, 0.01, 0.01
max values : 1.00, 1.00, 1.00, 1.00 Multi-Layer Rasters
Calling plot() on a multi-layer raster shows us one sub-plot per layer
Multi-Layer Rasters
Task: Calculate the mean habitat suitability around FL waters
class : SpatRaster
size : 300, 720, 4 (nrow, ncol, nlyr)
resolution : 0.5, 0.5 (x, y)
extent : -180, 180, -75, 75 (xmin, xmax, ymin, ymax)
coord. ref. : lon/lat WGS 84 (EPSG:4326)
source(s) : memory
names : Caretta_caretta, Chelonia_mydas, Dermoch~oriacea, Eretmoc~bricata
min values : 0.51, 0.51, 0.55, 0.51
max values : 1.00, 1.00, 1.00, 1.00 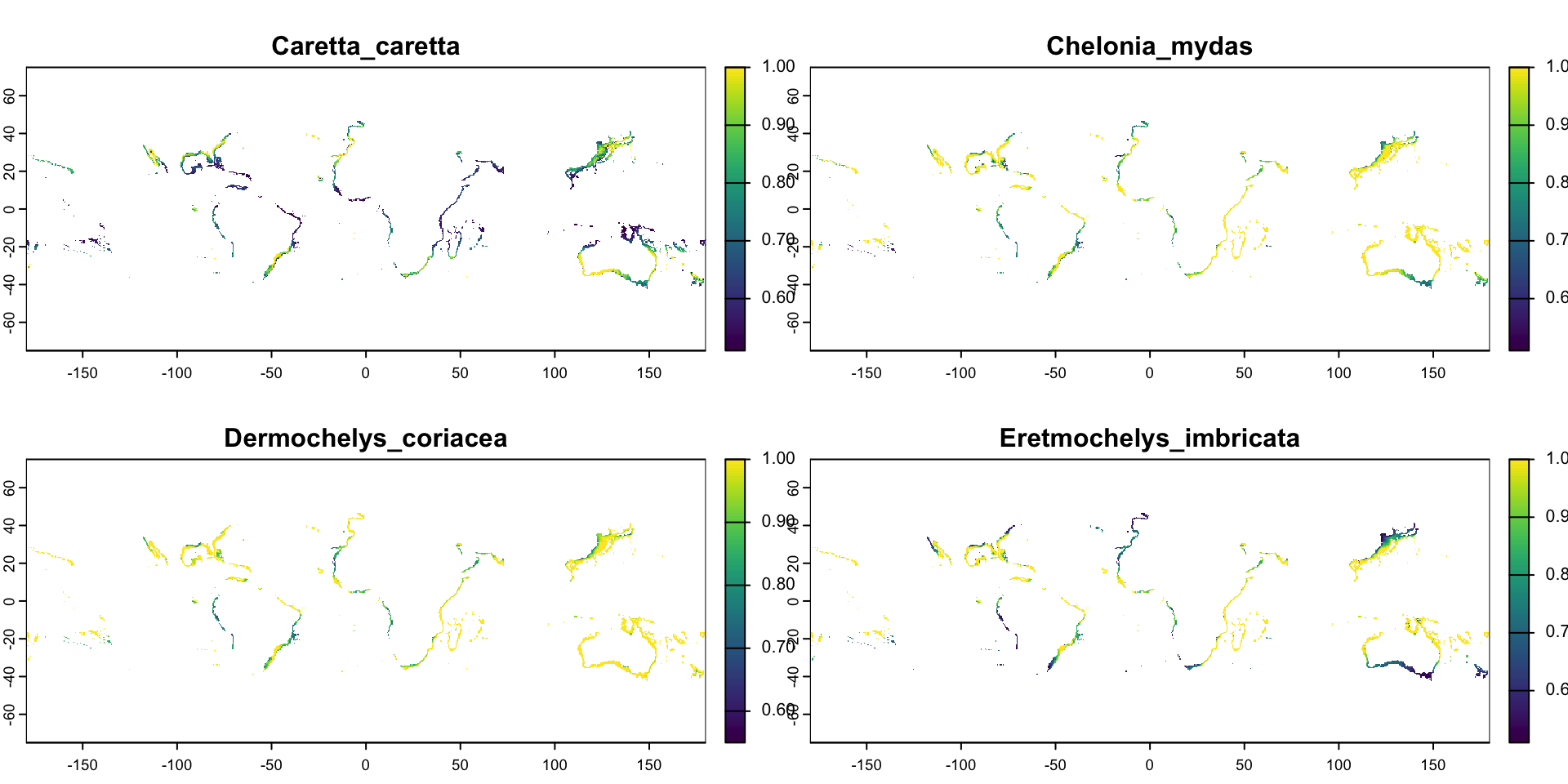
Multi-Layer Rasters
Task: Calculate the mean habitat suitability around FL waters
turtle_habitat <- turtles |> # Start from turtles
# And pipe into filter()
filter(Caretta_caretta > 0.5,
Chelonia_mydas > 0.5,
Dermochelys_coriacea > 0.5,
Eretmochelys_imbricata > 0.5) |>
# Then crop it to FL counties
crop(st_buffer(FL_counties, dist = 100000)) |> # Any guesses on what st_buffer is doing?
mean()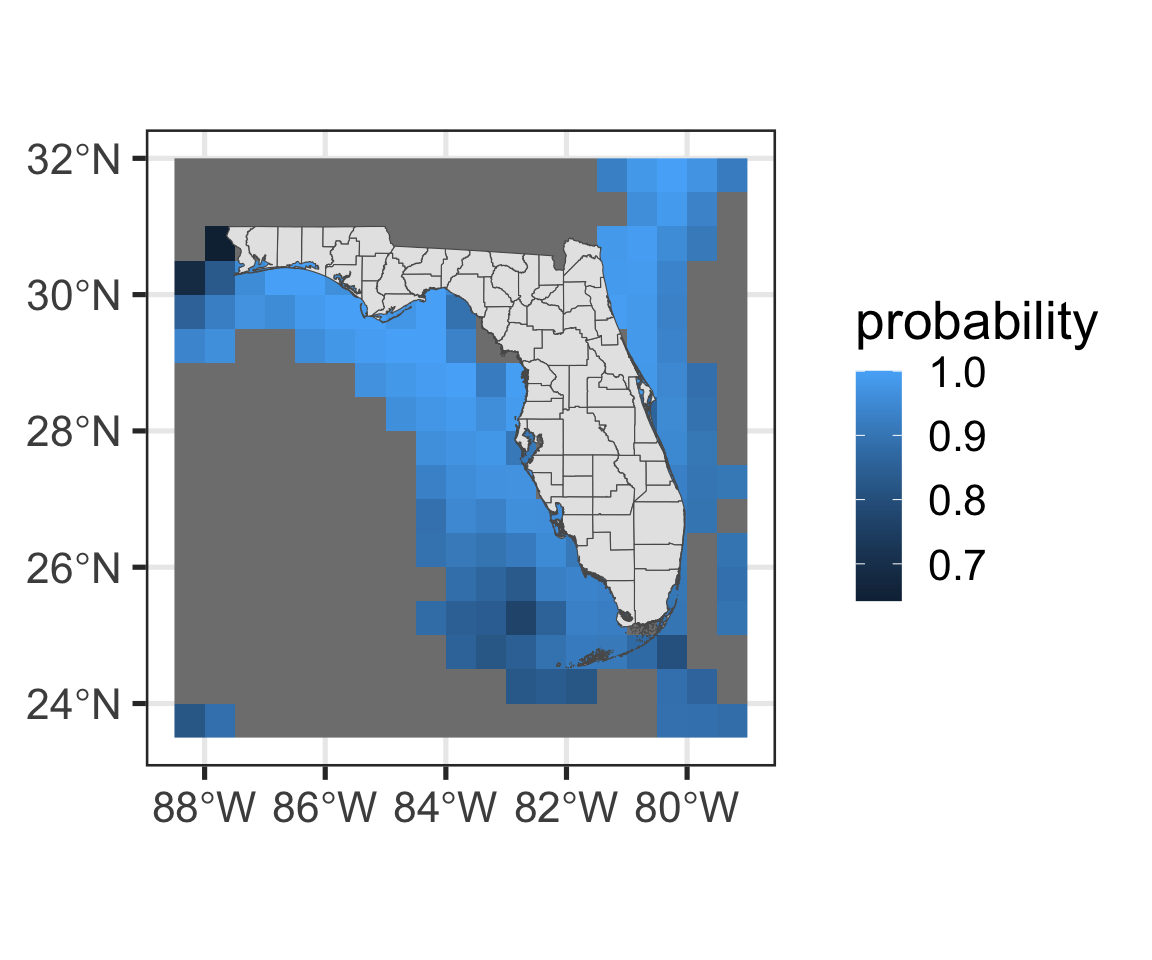
Wrap Up
- Raster Data Model: Continuous grid representation with pixels/cells of uniform size
- Ideal for continuous data (elevation, temperature, population density)
- More storage-efficient than vector data
- Each cell holds a single value per layer
- The
terraPackage:- Modern replacement for
{raster}package - Use
rast()to read raster files (.tif,.tiff,.asc,.nc)
- Modern replacement for
- The
tidyterraPackage:- Extends
terrawith tidyverse-friendly functions - Use
terrafor spatial operations (e.g.,crop()) - Use
tidyterrafor attribute operations (e.g.,filter(),mutate())
- Extends
- Multi-Layer Rasters:
- Combine multiple rasters with
c() - All layers must have same extent and resolution
- Use
mean(),sum(), etc. to aggregate across layers
- Combine multiple rasters with
Before Next Class:
- Update RStudio
- Install
terra,tidyterra, andggspatial